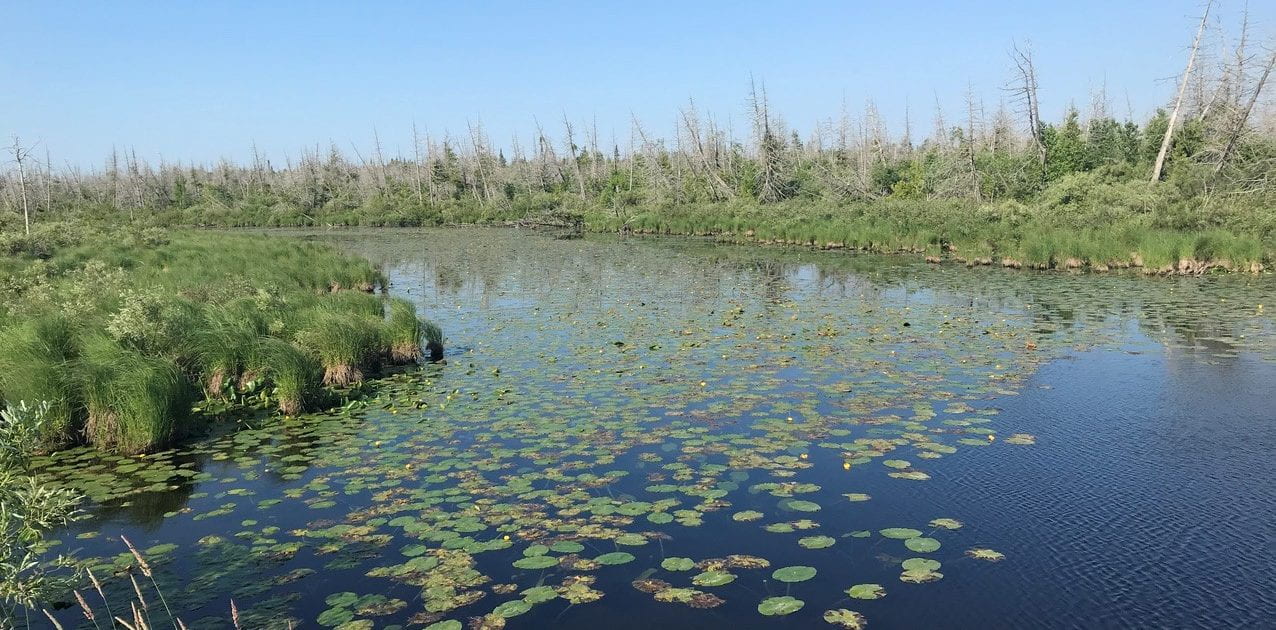
Evolution occurs in heterogeneous environments, over large geographic ranges, and in the presence of variable species assemblages. Our research focuses on how genetic variation is organized both at the species level and in populations of each species, what evolutionary processes regulate genetic variation, and how ecology can affect these evolutionary processes.
Current Projects
Sex determination and hybridization in Catostomus suckers
In Catostomus fishes (suckers) in the Upper Colorado River basin (Colorado, Wyoming, and Utah, USA), complex hybridization dynamics involving six different parental species lead to extremely variable genomic outcomes of hybridization. Hybrids are more constrained in some crosses than others, but it is unclear what mechanisms drive reproductive isolation. We aim to 1) identify regions of the genome associated with sex determination in Catostomus suckers, and 2) apply this knowledge to better describe hybridization outcomes.
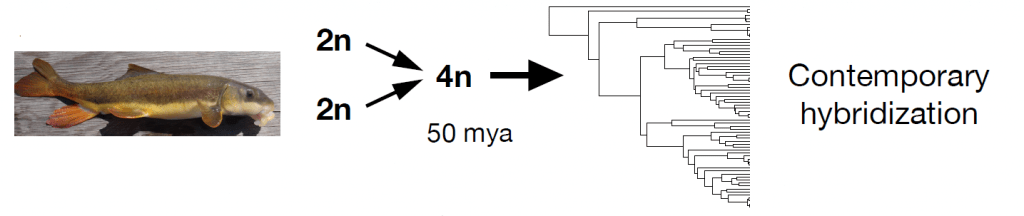
Above: Catostomus species are descended from an allotetraploid ancestor and experience contemporary hybridization
Sex determination and hybridization in Chrosomus dace
All-female, asexually reproducing populations of dace can be found across the US and Canada. It is believed that these lineages originated from hybridization events between Chrosomus eos and C. neogaeus, but the origins of these lineages are still poorly understood. No contemporary hybridization is known to occur. We aim to identify the history of hybridization and any potential backcrossing, as well as comparing sex determination mechanisms in parental lineages to identify potential incompatibilities.
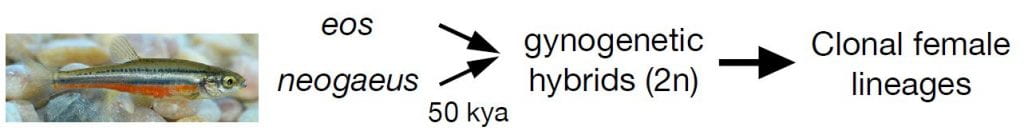
Above: Chrosomus eos and C. neogaeus hybridized to produce all-female clonal lineages approximately 50,000 years ago
Genetic structure of fish populations in agricultural streams
As a part of the University of Guelph’s Food From Thought program, and in collaboration with several other groups in the Integrative Biology department, this project aims to determine the effects of agricultural land use and habitat fragmentation on the genetic structure of fish populations. We are comparing genetic differentiation and genetic diversity across several species sampled at these sites, including creek chub and common shiner. We will also identify any hybridization that has occurred; common shiner is known to hybridize with several other species found at these sites, and hybridization is often more common in disturbed landscapes.
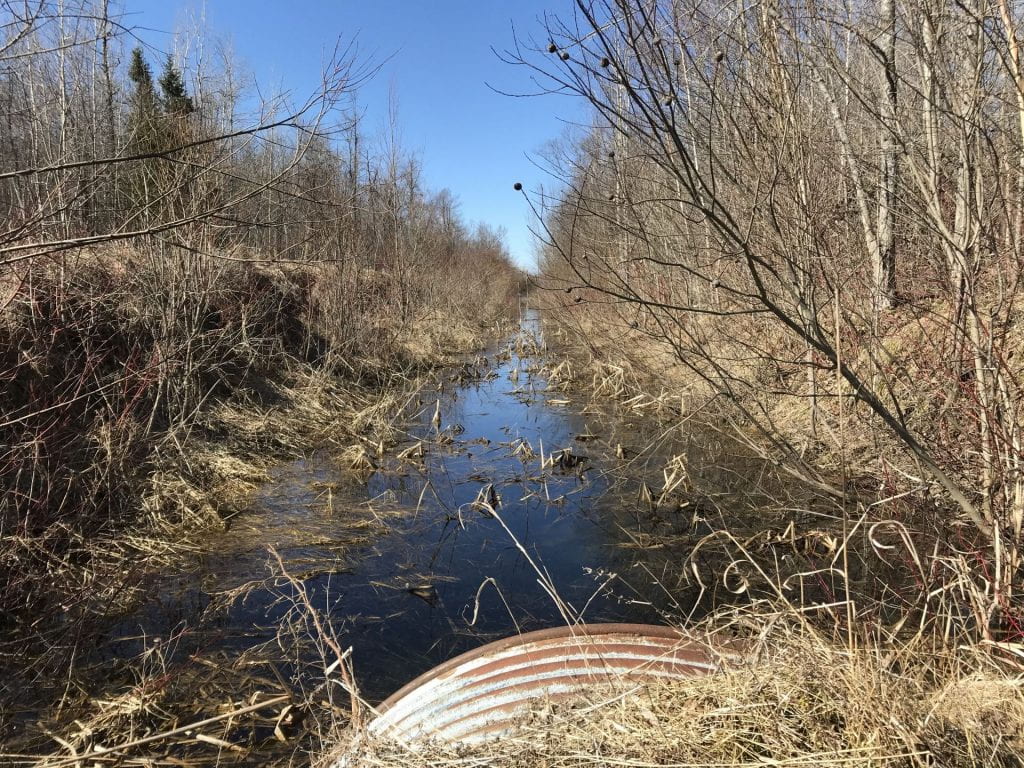

Above: An agricultural waterway in Southern Ontario
Efficacy of a Resistance Board Weir intervention to reduce Catostomus hybridization
Native and non-native Catostomus suckers are known to hybridize extensively in the Gunnison River Basin located in Colorado. It is thought that maladaptive hybrids are causing demographic swamping in this population, resulting in the potential loss of native species. Implementing an intervention approach such as a Resistance Board Weir (RBW) to stop passage of non-native suckers provides a plausible solution to this hybridization problem. Using genetic evidence, we aim to understand how successful an RBW intervention is as a means to reducing hybridization and how ancestry trends of larval offspring vary across the spawning habitat.

Above: A RBW installed in the study site, Roubideau Creek, Colorado.
Machine learning to identify fish hybrids
Liz Mandeville Sam Arevalo Amy Pitura
Rainbow trout (Oncorhynchus mykiss) have been widely introduced across the world, and can hybridize with native trout. In the North Fork Shoshone River drainage (Cody, Wyoming), rainbow trout hybridize with native Yellowstone cutthroat trout (Oncorhynchus clarki bouvieri). The resulting parental and hybrid juveniles can be difficult to identify without genetic analysis. Machine learning for analysis of images might provide a useful tool to identify phenotypes of these genotypes. We are training a model using images of previously sequenced individuals – i.e., genetic identity is known – as labeled training data. If successful, this approach could serve as a useful tool in conservation for identifying fish species.
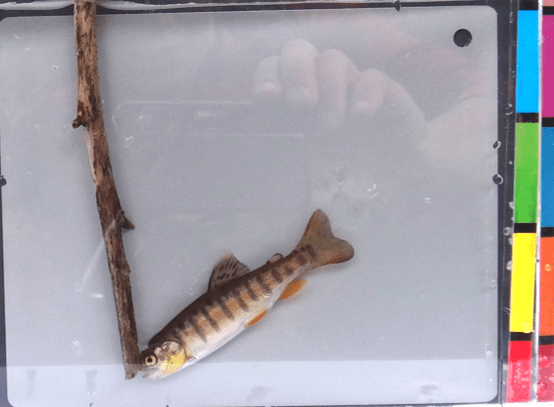

Above: Example photo of a juvenile trout used to train and test machine learning models






